Current Research
We aim to understand the interplay between chromatin and RNA in the network of gene regulatory processes. The lab focuses on three main questions:
1) How the piRNA pathway – a small non-coding RNA pathway – regulates transcription and chromatin structure.
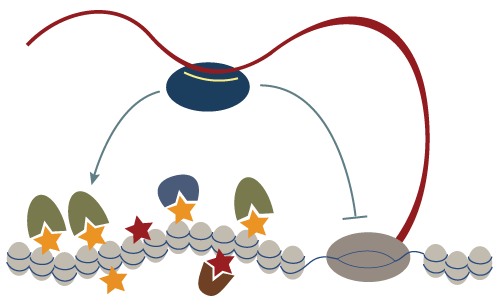
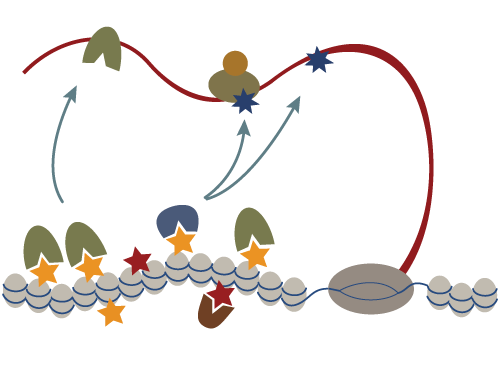
Chromatin structure plays a key role in regulating expression of the underlying genomic sequence. Yet we know little about the molecular basis of establishing diverse chromatin states on different genomic regions. Non-coding RNAs are emerging as key regulators of gene expression and are crucial for survival, development and differentiation. They can serve to convey sequence-specificity to regulatory processes. In animals, small RNAs were mostly associated with post-transcriptional regulation through a phenomenon termed RNA interference or RNAi. Our recent studies have revealed that a germline specific small RNA pathway, the piRNA pathway, also acts to regulate transcription by directing establishment of a repressive chromatin over its genomic targets. Our goal is to elucidate the exact mechanism through which the piRNA pathway induces transcriptional repression by identifying key factors in this silencing process and by elucidating their functions.
2) How piRNA serve as transgenerational epigenetic signal to establish gene expression patterns in the progeny.
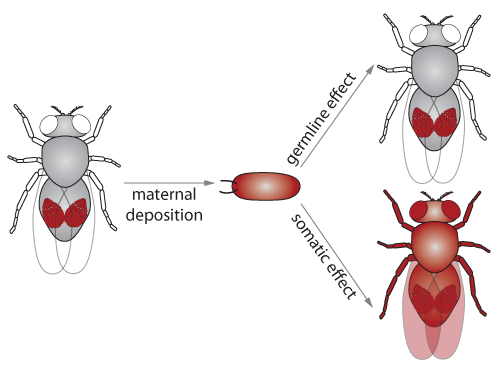
In Drosophila, piRNAs and the associated Piwi proteins accumulate during germline development and are deposited by the mother into the egg. Accordingly, piRNAs are present in the progeny at early stages of development and can impact chromatin state of its targets. Although the piRNA pathway is largely restricted to the germline, this presence in the early embryo results in exposure of the soma allowing the pathway to impact transcriptional profiles of somatic tissues as well. We are investigating how maternally inherited piRNAs serve as a transgenerational epigenetic signal to establish expression patterns during the development of the progeny.
3) How chromatin impacts post-transcriptional RNA fates.
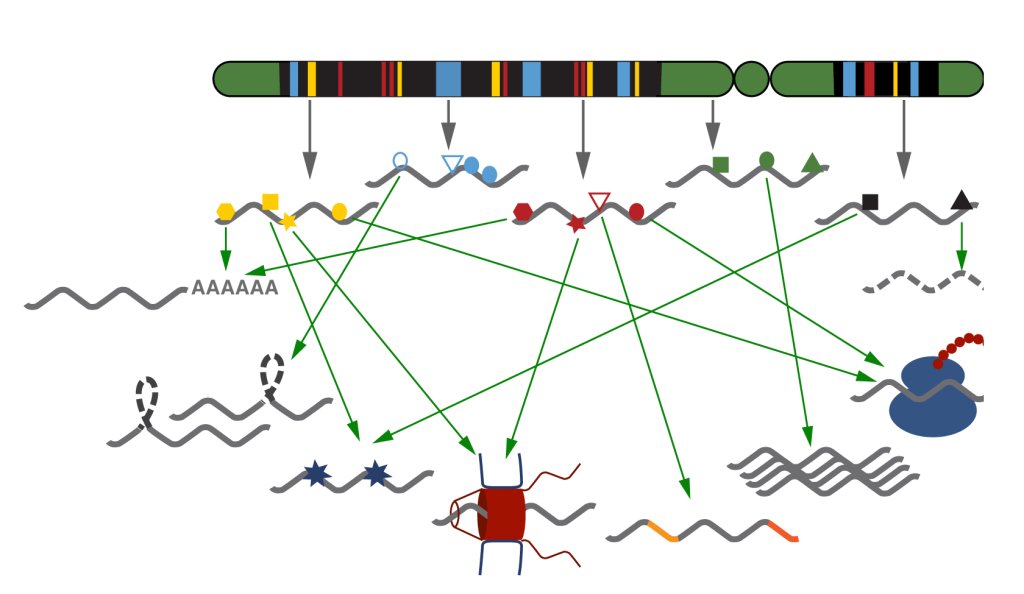
Chromatin is known to impact expression of the underlying genomic sequence. Regulation of transcription and the control of the post-transcriptional fate of RNAs – such as RNA processing, RNA editing, nuclear export, translation and RNA degradation – are often viewed as two independent processes. However, accumulating evidence suggests that the two steps are tightly linked and that chromatin is also involved in post-transcriptional gene regulation: some proteins that define the future fate of an RNA bind co-transcriptionally in a manner that depends on specific transcription factors and chromatin structure of the locus. We use a systems biology approach to investigate how chromatin influences the fate of emerging transcripts.